Patient QA System Development with RAG
Building an accurate, fluent patient question-answering system using Retrieval-Augmented Generation
Project Overview
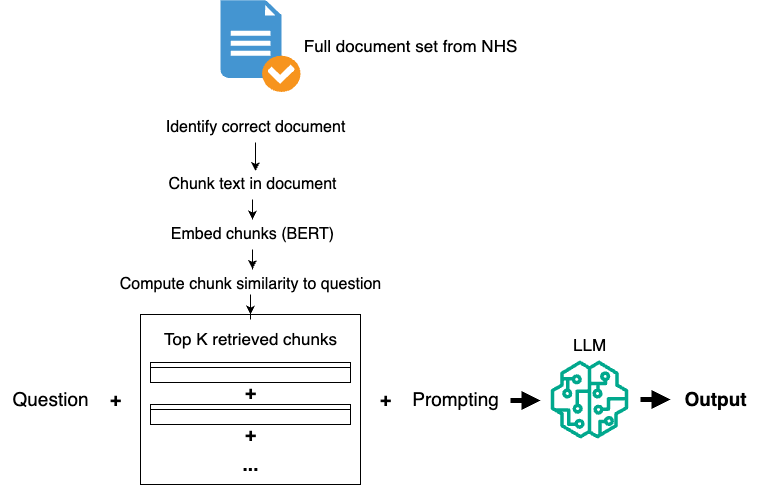
Patient QA systems provide health-related information and guidance to the public, requiring strict safeguards to ensure accuracy and reliability. This project developed a Retrieval-Augmented Generation (RAG) pipeline that retrieves vetted NHS documents and generates accurate, fluent responses to patient health questions.
Challenge: Traditional patient QA systems rely on retrieval-only methods to avoid hallucinations, sacrificing fluency and conversational quality. This project aimed to achieve both high accuracy and natural language fluency using advanced RAG techniques.
Problem Statement
Existing patient QA chatbots (e.g., WHO COVID-19 WhatsApp bot, NHS-Alexa) use keyword matching and simple retrieval methods that return rigid, template-based responses. While safe from hallucinations, they lack conversational fluency. Academic prototypes using generation models often lack evaluation metrics or fail to use vetted health authority documents, making them unsuitable for deployment.
Research Gap: No published study had demonstrated an auditable RAG pipeline that:
- Grounds every response in NHS source documents
- Achieves very high factual precision
- Maintains natural, fluent conversational tone
- Provides transparent, reproducible evaluation metrics
Approach & Methodology
Dataset
The project used the OpenGPT dataset (CogStack, UCL/KCL), containing 211 expert-validated question-answer pairs across 23 diseases, each linked to NHS reference documents. This enabled rigorous evaluation of retrieval and generation quality against gold-standard medical content.
Technical Architecture
Two systems were developed:
1. Baseline Model
- Retrieval: TF-IDF sparse retrieval (k=5 chunks)
- Chunking: Paragraph-based splitting on double newlines
- Generation: Phi-4-mini-instruct LLM with basic prompting
2. Optimized Model
- Retrieval: BioBERT dense embeddings + FAISS vector search
- Chunking: Intelligent merging (minimum 10 words per chunk)
- Top-k: Increased from 5 to 15 retrieved chunks
- Prompting: Zero-shot instruction tuning for conciseness and fidelity
- Generation: Phi-4-mini-instruct with constrained generation
Key Innovations
1. Dense Retrieval with BioBERT
BioBERT embeddings capture both contextual and biomedical meaning from sub-word tokenization, enabling semantic matching between lay patient questions and clinical NHS text. Combined with FAISS for efficient vector search, this approach significantly improved retrieval relevance over keyword-based TF-IDF.
2. Improved Text Chunking
Analysis revealed that naive paragraph splitting created many meaningless short chunks (e.g., "NHS", "Skip to main content"). The optimized chunking method merged adjacent chunks shorter than 10 words, reducing average chunks from 21.1 to 12.5 per document while improving semantic coherence.
3. Prompt Engineering
System prompts were refined based on analysis showing reference answers averaged 1.97 sentences. The optimized prompt instructed the model to provide "1-2 complete sentences unless the question requires elaboration," improving conciseness and alignment with expert answers.
Results & Impact
Manual Evaluation (50-question benchmark)
- Omission: "No omission" scores improved from 62% to 68%
- Hallucination: Optimized model achieved 100% accuracy (baseline: 2 minor errors)
- Fluency: Both models achieved 100% "perfectly fluent" ratings
Key Finding: The optimized RAG pipeline achieved a 40% improvement in ROUGE-Lsum precision (0.39 → 0.55) while maintaining perfect fluency and eliminating all hallucinations—demonstrating that accuracy and conversational quality are not mutually exclusive in patient QA systems.
Technical Insights
Retrieval Method Comparison
Dense retrieval with BioBERT outperformed TF-IDF particularly at higher k values (k=15), as the semantic understanding allowed better ranking of relevant chunks even when vocabulary didn't exactly match between questions and NHS text.
Top-k Optimization
Increasing k from 5 to 15 yielded the largest single improvement in ROUGE-Lsum precision (+18.3%), suggesting that providing the LLM with more context enabled better answer synthesis while maintaining accuracy.
Limitations & Future Work
- Some omissions of critical escalation information (e.g., "when to seek emergency care")
- Chunking could be further improved with sliding-window approaches (128-2048 tokens with overlap)
- Potential for automated hyperparameter optimization using Bayesian methods ("AutoRAG")
- Future work: framework for context-dependent chunking strategies
Tools & Technologies
- BioBERT: Biomedical language model for contextual embeddings
- FAISS: Efficient vector similarity search
- Phi-4-mini-instruct: Microsoft's instruction-tuned LLM
- ROUGE & BERTScore: Evaluation metrics for text generation quality
- Python, PyTorch, Transformers: Implementation stack
Significance
This project demonstrates that Retrieval-Augmented Generation can produce patient-facing health information that is simultaneously:
- Safe: Zero hallucinations, fully grounded in vetted NHS documents
- Accurate: 92.6% BERT F1 semantic similarity to expert answers
- Fluent: 100% natural, conversational responses
- Transparent: Auditable retrieval pipeline with comprehensive metrics
By publishing detailed methodology and evaluation metrics, this work addresses a critical gap in patient QA system research and provides a foundation for deploying trustworthy AI-powered health information systems.